RoBiC System Overview
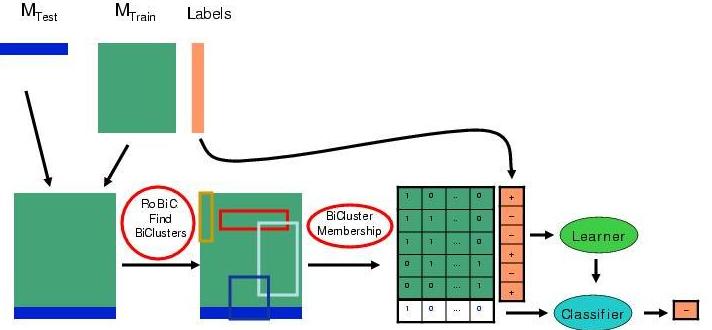
The RoBiC system learns microarray classifiers by first
reducing the dimentionality of data matrix using biclusters.
In general, a bicluster is a subset of genes and a subset of samples whose
expression values have similar patterns;
here, each bicluster is a sparse rank-one matrix -- ie, the outer product of
two sparse vector.
TechReport (9page): "Using Rank-1 Biclusters to
Classify Microarray Data" (9/Apr/07)
TechReport (7page): "Using Rank-One Biclusters to
Classify Microarray Data" (21/May/07)
Nasimeh
Asgarian,
Russell Greiner
MSc Dissertation: "Using Rank-1 BiClusters",
(Jan 2007; Nasimeh Asgarian)
Related Algorithms and Approaches
- Approaches related to RoBiC:
*
How RoBiC differs from SVD and related approaches
* How RoBiC relates to other relevant results.
- RoBiC finds a set of BiClusters, then uses them to produce a classifier.
This page presents
other ways to use BiClusters to build a classifier
- RoBiC uses a particular "hinge" function to decide which patients and
which genes belong to a bicluster.
This page describes
other "hinge functions"
- As suggested by the Figure above, we first form biclusters based on both
test and (unlabeled) training instances.
This page describes how this compares
to simply adding a single test instance at a time (to the training set)
when finding the biclusters.
Details of Empirical Studies
Request
- If you have better results on these data sets, based on hold-out
data (or CrossValidation), please email the relevant information
- We are looking for other microarray datasets:
- Binary classification labels
(either included with each sample, or better:
some withheld until we produce our predictions)
- Complete data (ie, include a meaningful numeric score for each gene/sample pair)
If you have such datasets, please send them to us.
We will, of course, accept standard confidentiality agreements;
just let us know.
Our system uses data in the same format as the Plaid System;
see here.
(Of course, we can convert from other formats.)
Reading Group

